Rocco Meli
Publications
Journal Articles
[11] M. Iannuzzi, J. Wilhelm, F. Stein, A. Bussy, H. Elgabarty, D. Golze, A.-S. Hehn, M. Graml, S. Marek, B. Sertcan Gökmen, C. Schran, H. Forbert, R. Z. Khaliullin, A. Kozhevnikov, M. Taillefumier, R. Meli, V. V. Rybkin, M. Brehm, R. Schade, O. Schütt, J. V. Pototschnig, H. Mirhosseini, A. Knüpfer, D. Marx, M. Krack, J. Hutter, T. D. Kühne, The CP2K Program Package Made Simple, J. Phys. Chem. B (2026)
[10] J. Biddiscombe, M. Simberg, A. Reverdell, R. Solcà, A. Invernizzi, R. Meli, J. Schuchart, On the integration of lightweight tasks with MPI using the C++ 26 std::execution “Senders” API, SC Workshops ‘25 (2025)
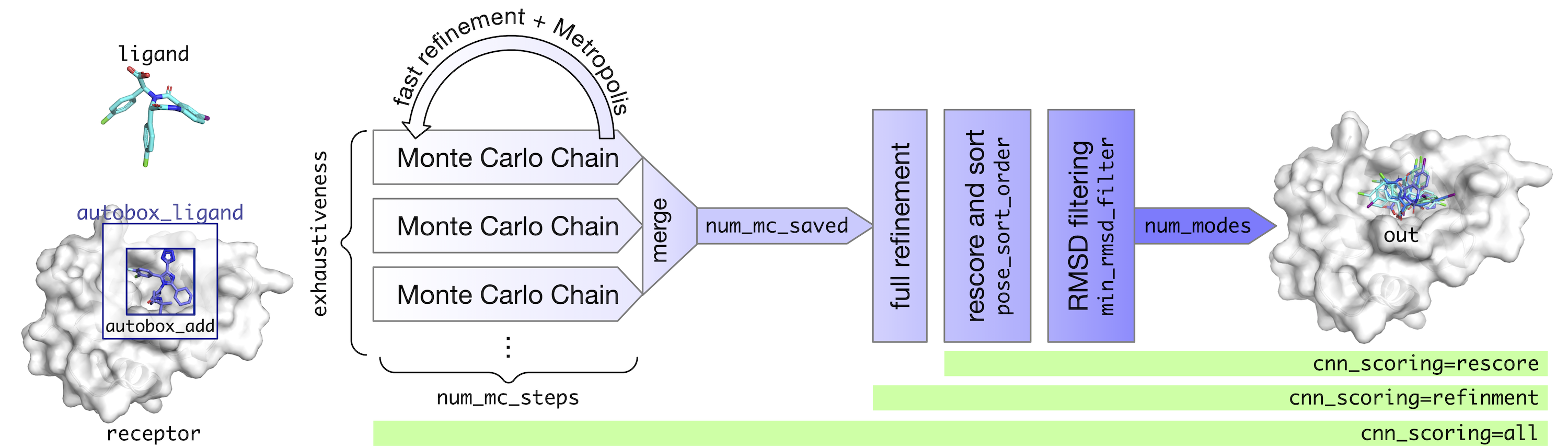
[9] A. T. McNutt, Y. Li, R. Meli, R. Aggarwal, D. R. Koes, GNINA 1.3: the next increment in molecular docking with deep learning, J. Cheminform. 17, 28 (2025)
[8] R. Solcà, M. Simberg, R. Meli, A. Invernizzi, A. Reverdell, J. Biddiscombe, DLA-Future: A Task-Based Linear Algebra Library Which Provides a GPU-Enabled Distributed Eigensolver, Asynchronous Many-Task Systems and Applications. WAMTA 2024, Springer, 14626 (2024) </a>
[7] R. Meli, G. M. Morris, P. C. Biggin, Scoring Functions for Protein-Ligand Binding Affinity Prediction Using Structure-based Deep Learning: A Review, Front. Bioinform., 2:885983 (2022)
[6] R. Meli, A. Anighoro, M. Bodkin, G. M. Morris, P. C. Biggin, Learning Protein-Ligand Binding Affinity with Atomic Environment Vectors, J. Cheminform. 13, 59 (2021)
[5] A. McNutt, P. Francoeur, R. Aggarwal, T. Masuda, R. Meli, M. Ragoza, J. Sunseri, D. R. Koes, GNINA 1.0: Molecular docking with deeplearning, J. Cheminform. 13, 43 (2021)
[4] R. Meli and P. C. Biggin, spyrmsd: symmetry-corrected RMSD calculations in Python, J. Cheminform. 12, 49 (2020)
[3] E. Liberatore, R. Meli and U. Rothlisberger, A Versatile Multiple Time Step Scheme for Efficient Ab Initio Molecular Dynamics Simulations, J. Chem. Theory Comput. 14, 2834-2842 (2018)
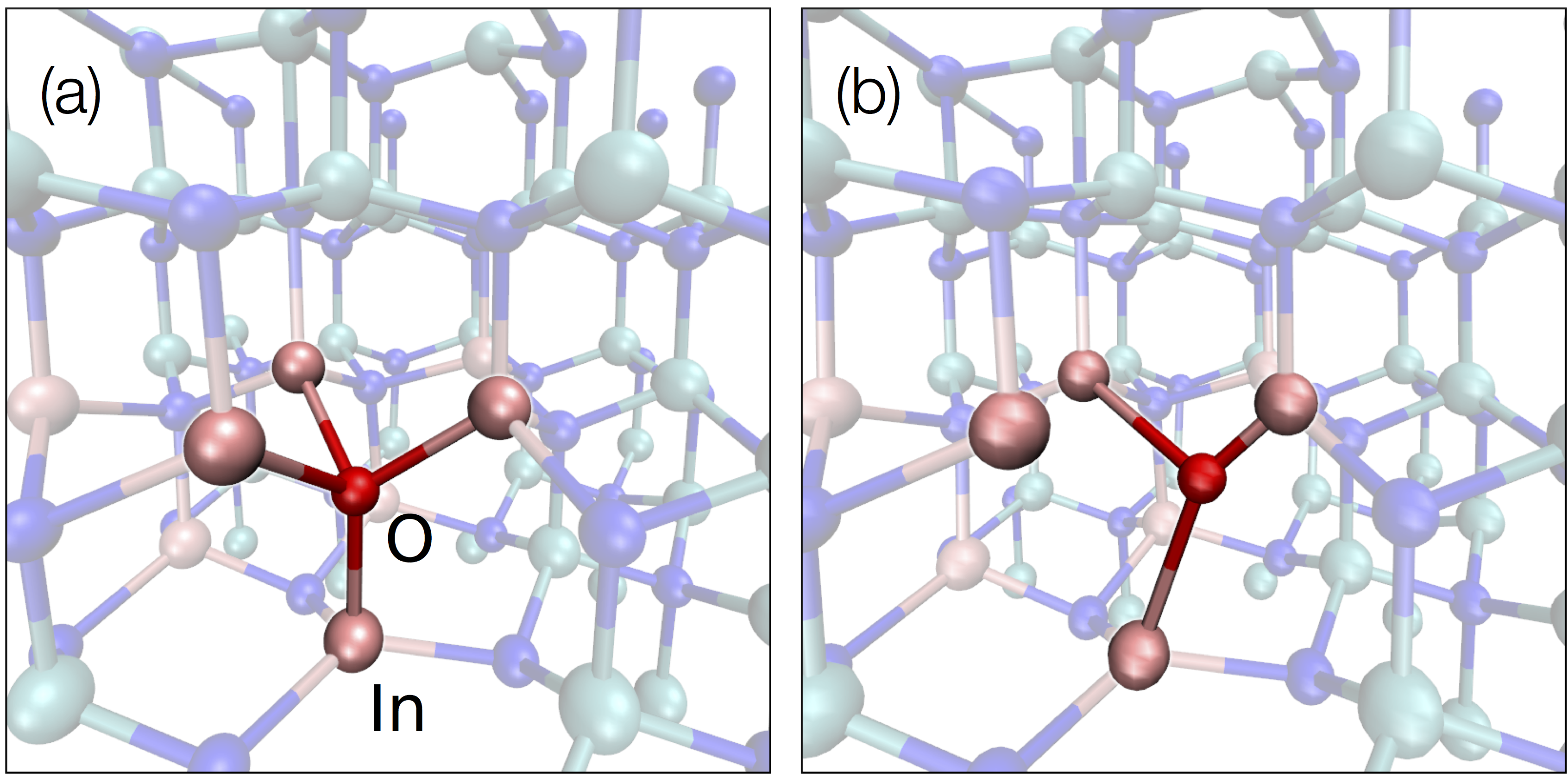
[2] R. Meli, G. Miceli and A. Pasquarello, Oxygen DX center in In0.17Al0.83N: Nonradiative recombination and persistent photoconductivity, Appl. Phys. Lett. 110, 072101 (2017)
[1] J. Beal, T. Haddock-Angelli, M. Gershater, K. de Mora, M. Lizarazo, J. Hollenhorst, R. Rettberg, iGEM Interlab Study Contributors, Reproducibility of Fluorescent Expression from Engineered Biological Constructs in E. coli, PLoS ONE, 11 (2016)
Preprints
[I] F. Manby, T. Miller, P. Bygrave, F. Ding, T. Dresselhaus, F. Batista-Romero, A. Buccheri, C. Bungey, S. Lee, R. Meli, K. Miyamoto, C. Steinmann, T. Tsuchiya, M. Welborn, T. Wiles, Z. Williams, entos: A Quantum Molecular Simulation Package, ChemRxiv (2019)